
Received: SeptemAccepted: DecemPublished: December 18, 2017Ĭopyright: © 2017 Arafa et al. PLoS ONE 12(12):Įditor: Mark Gijzen, Agriculture and Agri-Food Canada, CANADA In addition, the approach developed in this study provides a model for identification of other genes for attractive agronomical traits.Ĭitation: Arafa RA, Rakha MT, Soliman NEK, Moussa OM, Kamel SM, Shirasawa K (2017) Rapid identification of candidate genes for resistance to tomato late blight disease using next-generation sequencing technologies. SNP and SSR markers linking to this region can be used in marker-assisted selection in future breeding programs for late blight disease, including introgression of new genetic loci from wild species. Among of them, two genes with missense mutations, Solyc06g071810.1 and Solyc06g083640.3, were considered to be potential candidates for disease resistance. Whole-genome resequencing analysis revealed that 298 genes in this region potentially had functional differences between the parental lines. Subsequent association analysis of genotypes and phenotypes of the mapping population revealed that a 6.8 Mb genome region on chromosome 6 was a candidate locus for disease resistance.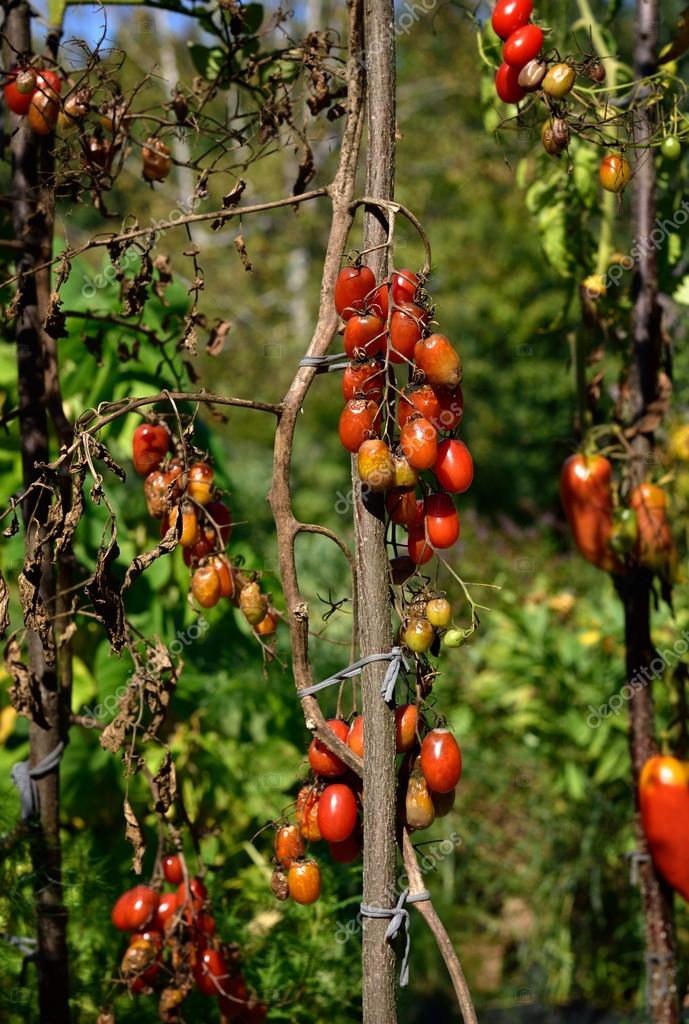 
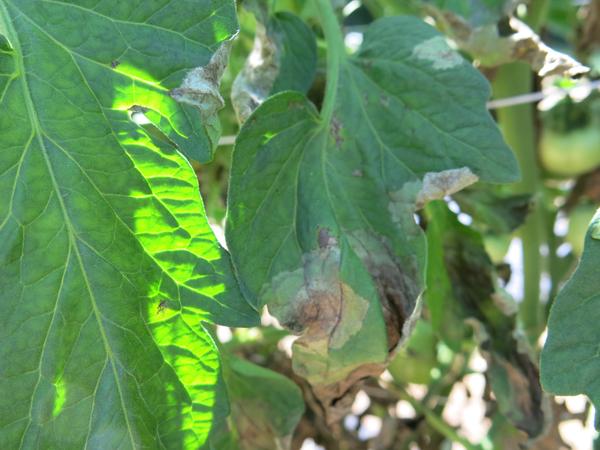
Using double-digest restriction site–associated DNA sequencing (ddRAD-Seq) technology, we determined 6,514 genome-wide SNP genotypes of an F 2 population derived from an interspecific cross. In the genome of a wild relative of tomato, Solanum habrochaites accession LA1777, we identified a new quantitative trait locus for resistance against blight caused by an aggressive Egyptian isolate of P. Wild relatives of tomato possess useful resistance genes against this disease, and could therefore be used in breeding to improve cultivated varieties. Tomato late blight caused by Phytophthora infestans (Mont.) de Bary, also known as the Irish famine pathogen, is one of the most destructive plant diseases.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
